WAVES
Summary
WAVES is a web application dedicated to bioinformatic tool integration. It provides an efficient way to implement a service for any bioinformatic software. Such services are automatically made available in three ways: web pages, web forms to include in remote websites, and a RESTful web services API to access remotely from applications. In order to fulfill the service’s computational needs, WAVES can perform computation on various resources and environments, such as Galaxy instances.
Availability and implementation
WAVES was developed with Django, a Python-based web framework. It was designed as a reusable web application. It is fully portable, as only a Python installation is required to run Django. It is licensed under GNU General Public License version 3.
WAVES components
WAVES-core
This is the main WAVES component.
- GitHub repository: https://github.com/lirmm/waves-core
- Documentation: http://waves-core.readthedocs.io
- Installation guide: http://waves-core.readthedocs.io/en/latest/installation.html
- Administration guide: http://waves-core.readthedocs.io/en/latest/user_doc/user_guide.html
- Tutorial to create a ‘Hello world’ service: http://waves-core.readthedocs.io/en/latest/user_doc/howto/howto.html
- Developer guide: http://waves-core.readthedocs.io/en/latest/dev_doc/dev_doc.html
- Python code examples to interact with WAVES API: http://waves-core.readthedocs.io/en/latest/dev_doc/sample_code.html
WAVES-Galaxy
This is a WAVES adapter dedicated to Galaxy. Using the BioBlend python library, this adapter enables Galaxy services import into WAVES-core. WAVES-Galaxy automatically recognizes the tools available in a Galaxy instance.
- GitHub repository: https://github.com/lirmm/waves-galaxy
- Documentation: http://waves-galaxy-adaptors.readthedocs.io
WAVES-demo
This is a custom WAVES instance for demo purpose. It was created to show what could be done with Django functionalities to custom a WAVES installation.
- GitHub repository: https://github.com/lirmm/waves-demo
- Documentation: http://waves-demo.readthedocs.io
- Demo: http://waves.demo.atgc-montpellier.fr
The changes that have been made are:
- Use a different skin for the back-office interface.
- Custom front-end interface.
- Override the Service class.
- Override the Authentication class.
Please note that anyone can login the back-office administration interface and create new services. Thus, for security reasons, we intentionally deactivated the jobs running procedure.
WAVES Singularity image
This is a Singularity container with a functional WAVES installation including two pre-configured services (‘Hello world’ and ‘PhyML’). For testing purpose.
- Singularity image: http://www.atgc-montpellier.fr/download/binaries/waves/wavestest.simg
- Documentation: http://waves-core.readthedocs.io/en/latest/extensions.html#just-have-a-try
Contact
You may contact the WAVES support by e-mail : waves [at] lirmm.fr
Other tools
PhyML 3.0
Overview: new algorithms, methods and utilities PhyML is a software package that uses modern statistical approaches to build phylogenetic trees from the analysis of alignments of nucleotide or amino acid sequences. The main tool in this package builds phylogenies under the maximum likelihood criterion. It implements a large number of substitution models coupled to efficient…

TFscope
Characterizing the binding preferences of transcription factors (TFs) in different cell types and conditions is key to understand how they orchestrate gene expression. TFscope is a machine learning approach that identifies sequence features explaining the binding differences observed between two ChIP-seq experiments targeting either the same TF in two conditions or two TFs with similar…
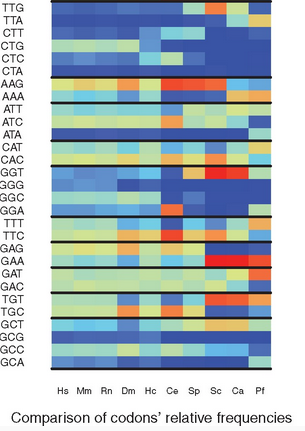
RSCU_RS: Measuring the bias in…
Overview Overview: In the protein coding sequences of a species, the 61 possible codons of the genetic code are not equally distributed. This observation is referred to as the Codon Usage Bias (CUB) of a species. Several measures have been proposed to quantify the CUB using the frequencies of codons in all RNA coding sequences…
